Contact Admission
International Collaboration
USTIN, Texas - Researchers from the University of Texas at Austin and the National Institutes of Health made a major breakthrough to the development of new coronavirus vaccines in 2019 by creating a scale map First 3D atom of a tuber part
Researchers at the University of Texas, Austin and the National Institutes of Health (NIH) created a 3D spike protein (P) ratio map of 2019-nCoV, a key breakthrough for development. vaccine COVID-19 disease for the new 2019 coronavirus infects human cells.
The coronaviruses have the genome of RNA, with a set size from 26 to 32,000 N base pairs (A-T, C-G). The structural proteins that encase the RNA genomes are: spike (S), envelope (E), membrane (M) and nucleocapsid (N)).
Spike protein S is an important part, the "key" to help coronaviruses attach and enter the cells of a specific host, in this case of disease COVID-19 the main host is human.
AUSTIN, Texas - Researchers from the University of Texas at Austin and the National Institutes of Health made a major breakthrough to the development of a new coronavirus vaccine in 2019 by creating a scale map The first 3D atom of a part of the virus. human cell infection.
Mapping this piece, known as the spike protein, is an essential step for researchers around the world to be able to develop vaccines and antivirals to fight the virus. The article was published on Wednesday, February 19 in Science journal.
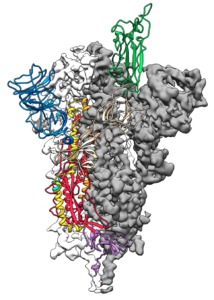
The science team is also working on a viable vaccine candidate derived from the study.
Jason McLellan, associate professor at UT Austin, head of research and his colleagues have spent many years researching other coronaviruses, including SARS-CoV and MERS-CoV. They developed methods to manipulate the coronavirus spike proteins into a shape that makes them easier to analyze and can effectively turn them into vaccine candidates. This experience has given them an advantage over other research groups working on new viruses.
As soon as we knew this was a coronavirus, we felt we had to learn about it, said Cameron McLellan, as we could be one of the first to have this structure. We know exactly what mutations are included here, because we have shown these mutations to be active against a wide range of other coronaviruses.
The majority of the research was done by the first study co-author, Ph.D. student Daniel Wrapp and research collaborator Nianshuang Wang, both at UT Austin.
Just two weeks after receiving the viral genome sequences from Chinese researchers, the team designed and produced samples of their steady spike proteins. It takes about 12 more days to reconstruct the 3D atomic ratio map, called molecular structure, the protein spike and submit the manuscript to Sciences, which conducts its peer review. Many of the steps involved in this process will usually take several months to complete.
Critical to its success is the cutting-edge technology called the cryo-EM microscope in UT Austin's new Sauer Laboratory. Cryo-EM allows researchers to create atom-scale 3D models of cellular, molecular and viral structures. We ended up being the first in part thanks to the infrastructure at Sauer Laboratories, says Martin McLellan. Software emphasizes the importance of funding basic research institutions. The molecule the team created and they had the structure that represented only the extracellular portion of the spike protein, but it was enough to induce an immune response in humans, and thus act as a vaccine. .
Next, McLellan's team plans to use the molecule to chase another line of attack against the virus that causes COVID-19, using the molecule as a user probe to isolate the produced antibodies. spontaneously from patients newly infected with coronavirus and with successful recovery. In sufficiently large amounts, these antibodies can help treat coronavirus infections soon after exposure. For example, antibodies that could protect military or health care workers are sent to an area with a high infection rate in too short a message for the vaccine immunity to take effect.

Other study co-authors are Kizzmekia Corbett and Olubukola Abiona at VRC; and Jory Goldsmith and Ching-Lin Hsieh at UT Austin.
Wang, Corbett, Graham and McLellan are inventors on a US patent application on the structure of the coronavirus mutant proteins in the prefabricated structure and their use in therapy. Wrapp, Wang, Corbett, Abiona, Graham, and McLellan are the inventors on the US patent application for the vaccine candidate described in this release.
This work is partly supported by the National Institutes of Health and the National Institute of Allergy and Infectious Diseases. The Sauer Structural Biology Laboratory is supported by the University of Texas at the Austin College of Natural Sciences, and the Texas Institute for Cancer Research and Prevention (CPRIT).
The University of Texas at Austin is committed to transparency and disclosure of all potential conflicts of interest. The university's investigator, who led the study, Jason McLellan, submitted the required financial disclosure forms to the university. McLellan holds the intellectual property rights that can generate revenue from the discovery described in this study.
Images and b-roll related to this study are available for download:
Other research
- PHAN ASIA UNIVERSITY OPEN EXPENSES RESEARCH COOPERATION WITH PASTEUR LILLE INSTITUTE- FRANCE ( 08:51 - 05/03/2020 )
- Importation and Human-to-Human Transmission of a Novel Coronavirus in Vietnam ( 14:57 - 31/01/2020 )
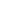
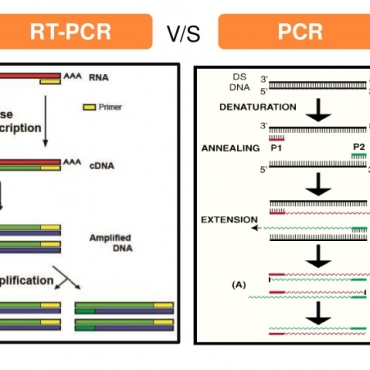
- PCR test in medical diagnosis ( 10:03 - 16/11/2019 )
- Announcement: Organizing Clinical Microbiology course ( 07:53 - 09/04/2019 )
- AI is so powerful that it can predict the moment of human death ( 08:42 - 01/04/2019 )
- World umbilical cord in Vietnamese hands ( 15:19 - 30/03/2019 )
- Stem cell magic ( 15:14 - 30/03/2019 )
- A new breakthrough in stem cell research ( 09:20 - 30/03/2019 )
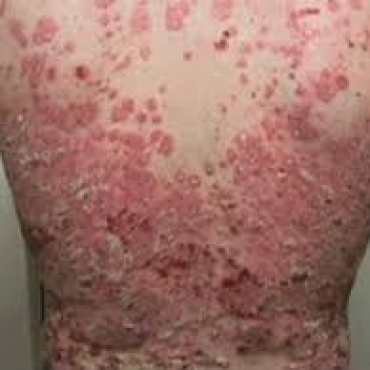
- Treatment of psoriasis ( 09:38 - 23/03/2018 )