Liên hệ tuyển sinh
Hợp tác Quốc tế
Evaluate the role of the microbiological molecular biology tests in the early detection of pathogens causing lower respiratory infections
BACKGROUND
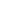
Based on the pathogenesis of the lower respiratory tract infections, the first specimen should be obtained from patients to detect the causative microbial agent is the sputum specimens. However, sputum is the contaminated specimen so that the big challenge must be overcome is to confirm the isolated bacteria is the pathogen, not the contaminated one. In addition, the most common pathogens of the lower respiratory tract infection are the fastidious bacteria requiring the immediate isolating on multiple media. Besides that, the previous using antibiotics are also affected on the culture’s results and the clinical lab could not culture the atypical bacteria and viruses. These are the obstacts that conventional culture method couldn’t overcome.
AIMS
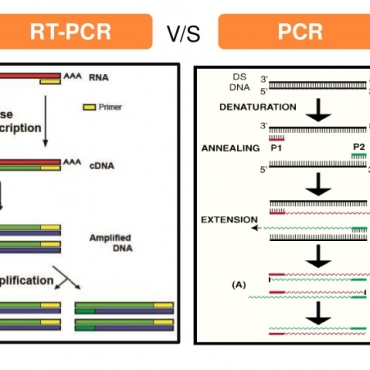
Evaluate the role of the multiplex real-time PCR that are currently apply in the clinical laboratory of Nam Khoa Company for detection of the micro-organism pathogens causing community acquired pneumonia (CAP) in children and in adults hospitalized in several hospitals in HCM city.
RESULTS - DISCUSSIONS
For children with CAP, one study on 52 cases with lobar pneumonia were analyzed. The results said that none of the sputum samples were positive with culture, and 100% of samples were detected pathogens by real-time PCR. The single pathogen were detected in 55.76% of cases, the multiple pathogens were detected in 44.24% of cases. The predominant pathogens detected were M. pneumoniae (69.2%), S. pneumoniae (55.8%). The other pathogens that were detected with lower ratio were H. influenzae (13.5%), M. catarrhalis (7.7%) and MRSA (3.9%). The real-time PCR results of this was demostrated in table 1
MATERIALS – METHODS
The samples were sputa (adult) or naso-tracheo aspirate sputa (children) collected from patients with CAP hospitalized in Children Hospital No.1 for children under 16YO, Cho Ray, Pham Ngoc Thach, Nhan Dan Gia Dinh and Can Tho Central Polyclinic Hospital for adult from 16YO. These samples were evaluated the reliable by Gram staining at the clinical lab of the hospital, the reliable samples were cultured there and then send to the clinical lab at Nam Khoa company for mul-tiplex real-time PCR detection of community bacteria, atypical bacteria, respiratory virus, and nosocomial bacteria. The real-time PCR that was carried-out at the clinical laboratory of Nam Khoa company was the STREAMLINE REAL-TIME PCR including the KingFISHER Flex instrument and the kit named NKDNARNAprep MAGBEAD that has been evaluated equivalent to Roche Magnapre system. The multiplex real-time PCR were prepared from Qiagen hot-start taq and the primers/taqman probes synthetized by Proligo Sigma (Singapore). The primers and the probes sequences were take from different published articles. For the RNA virus, before the real-time PCR, the cDNA synthesis were required with universal cDNA synthesis reagents from Roche. The criteria to define the pathogen was over 105 copies/ml for bacteria and over 102/ml for virus and atypical bacteria.
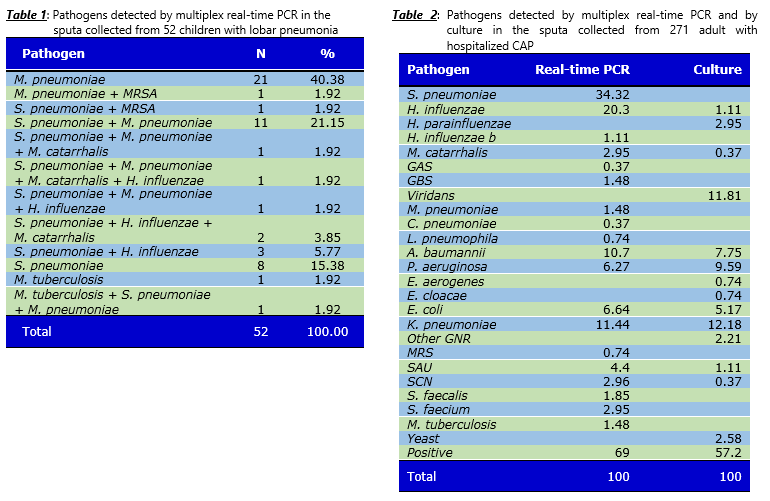
For adult with CAP, one study on 271 cases hospiatalized Cho Ray, Nhan Dan Gia Dinh, Pham Ngoc Thach, and Can Tho Central polyclinic hospitals were analysed. The received results said that 57.20% of the samples were [+] with culture and the predominant cultured pathogens were K. pneumoniae (12.18%), viridans streptococci (11.81%), P. aeruginosa (9.59%) and A. baumannii (7.75%). None of S. pneumoniae were detected and only 3 cases (1.11%) were cultured H. influenzae. By multiplex real-time PCR, 69% of cases were detected the pathogens and the predominant detected pathogens were S. pneumoniae (34.32%) and H. influenzae (20.3%), while the detection ratio of K. pneumoniae was lower (11.44%) as well as A. baumannii (10.7%), E. coli (6.64%) and P. aeruginosa (6.27%). The real-time PCR results and the culture results of this study was demonstrated in graph 2.
In this study, real-time PCR detected single pathogen in 40.22% and multiple pathogens in 28.71% of the CAP cases. There were some reasons to explain the different in the result of real-time PCR versus real-time PCR: (1) Viridans streptococci and H. parainfluenzae are considered as normal flora from the throat then its specific primer and probe were not included in the real-time PCR. (2) No specific primers and probes for E. aerogenes, E. cloacae and yeast in the real-time PCR. (3) Because of the previous using of antibiotic then the culture could not yield the S. pneumoniae and H. influenzae.
CONCLUSION
With the implement of the real-time PCR, the results of detection microbiological pathogens causing lower respiratory infection can arrive to the physicians timely, avoid the using of the empirical antibiotic treatment for longtime since the targeted antibiotic treatment can be done to the patients sort time after the clinical diagnosis.
Các tin khác
- Chẩn Đoán Quá Mức: Khi Ranh Giới Bệnh Tật Bị Làm Mờ ( 08:22 - 08/07/2025 )
- Nghiên cứu mới: Kiến thức liên tục của bác sĩ nội khoa ảnh hưởng đến chất lượng điều trị ( 10:56 - 19/06/2025 )
- Đặc điểm lâm sàng và cận lâm sàng bệnh nhân hậu COVID-19 tại bệnh viện đa khoa Tâm Trí Sài Gòn: một nghiên cứu hồi cứu cắt ngang ( 14:59 - 10/05/2022 )
- HIỆU QUẢ CỦA AZITHROMYCIN UỐNG SO VỚI GIẢ DƯỢC TRÊN CÁC TRIỆU CHỨNG COVID-19 Ở BỆNH NHÂN NGOẠI TRÚ VỚI NHIỄM SAR-COV-2- MỘT THỬ NGHIỆM LÂM SÀNG NGẪU NHIÊN ( 09:50 - 12/08/2021 )
- MỐI LIÊN QUAN GIỮA MẸ BỊ NHIỄM SARS-CoV-2 VÀ DỰ HẬU TRÊN TRẺ SƠ SINH ( 13:33 - 31/05/2021 )
- TẦM SOÁT UNG THƯ ĐẠI TRỰC TRÀNG ( 09:00 - 20/05/2021 )
- KHÁNG THỂ ĐẶC HIỆU SARS-COV-2 TRONG SỮA MẸ SAU CHỦNG NGỪA VACCINE COVID-19 Ở NHỮNG PHỤ NỮ ĐANG CHO CON BÚ ( 08:27 - 20/05/2021 )
- Lợi ích của việc Tránh Chăm sóc Y tế Không cần thiết Trong Đại dịch COVID-19 ( 14:01 - 15/05/2021 )
- Rapid and sensitive diagnostic procedure for multiple detection of pandemic Coronaviridae family members SARS-CoV-2, SARS-CoV, MERS-CoV and HCoV: a translational research and cooperation between the Phan Chau Trinh University in Vietnam and Universit ( 15:42 - 29/06/2020 )
- Các nghiên cứu của PCTU tại hội nghị quốc tế ISSAR 2019 - Gyeongju, Hàn Quốc ( 16:17 - 04/11/2019 )